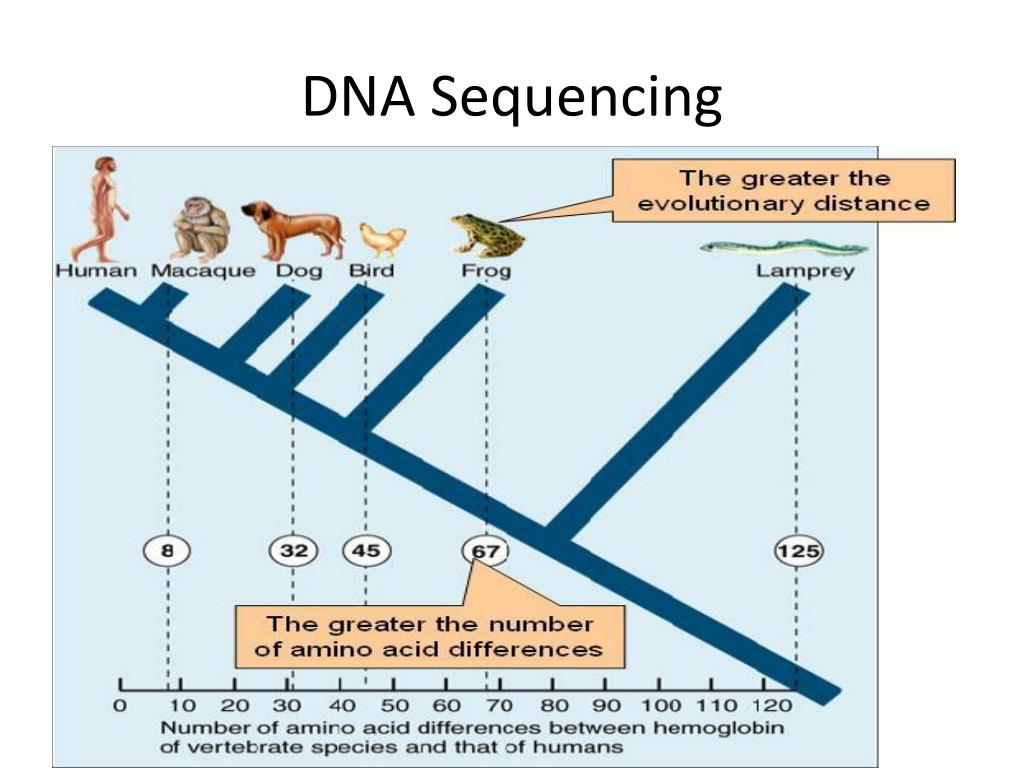
Clustal Omega – an alignment program that compares multiple sequences of DNA.GenBank – a genetic database that serves as an annotated collection of DNA sequences.Gene sequences from different species can be identified and then compared using two online resources: If you and another person both have the same ancestor, there’s a chance that you both inherited some of the same DNA. Since every person inherited DNA from their parents, who inherited it from their parents, and so on, a person’s DNA is made up of the DNA of their ancestors. We share more genes with organisms that are more closely related to us. The percentage of genes or DNA that organisms share records their similarities. What is the reason for DNA similarity?ĭue to billions of years of evolution, humans share genes with all living organisms. That’s because closely related species most likely diverged from one another fairly recently in the evolutionary span. Similarity analysis of DNA sequence is a process to compare unknown DNA sequences with known ones for inferring the functions of unknown ones it also can provide evolutionary information of the same gene sequence in different species, which is basic approach to comprehend the biological information in DNA sequence … What does it indicate if two organisms have similar DNA sequences?īecause the DNA sequence determines a protein’s amino acid sequence, a gene shared by two closely related organisms should have similar, or even identical, amino acid sequences. 9 What information can be obtained from DNA sequence?. 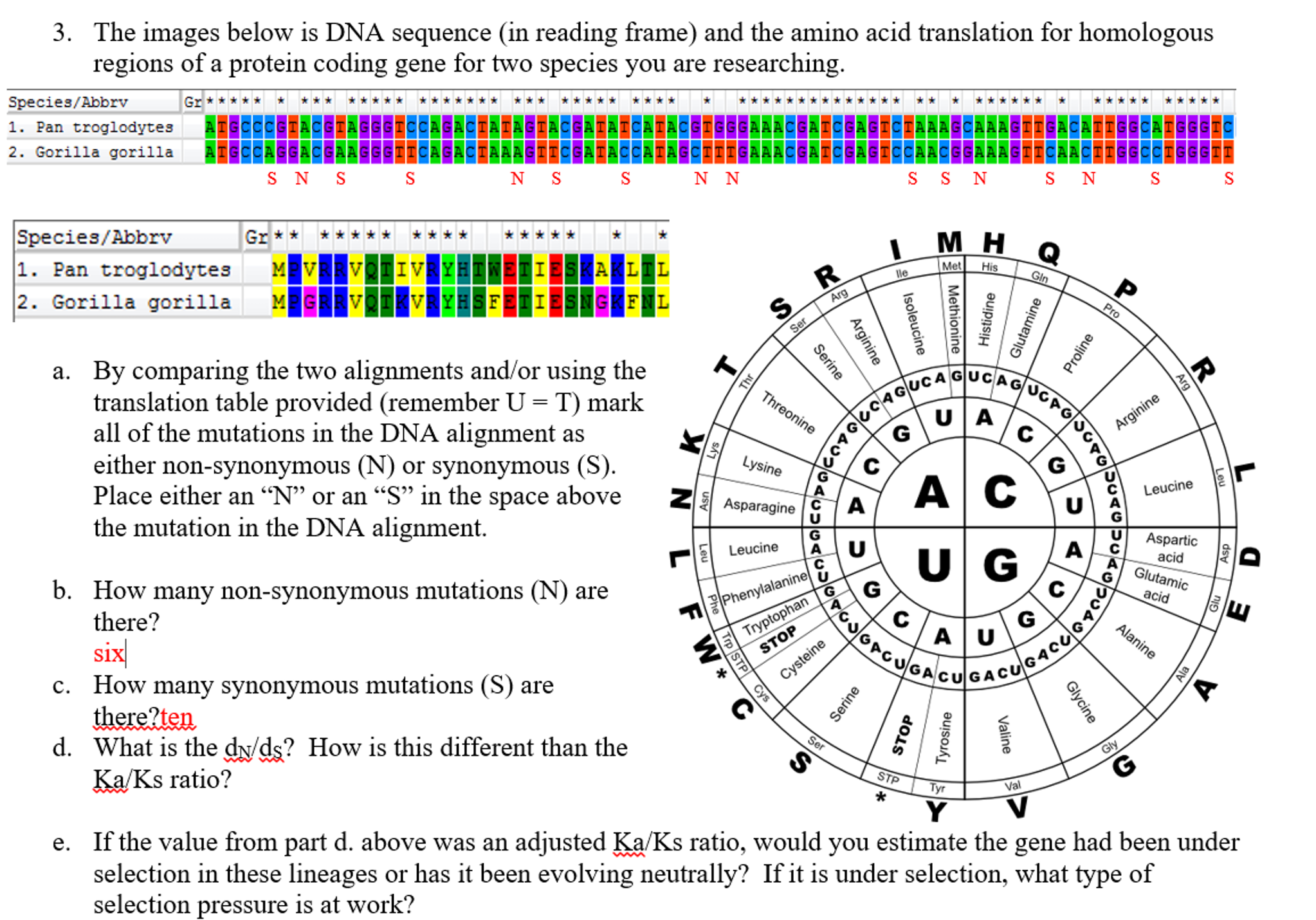
8 How do scientists study gene sequences?.6 How does DNA compare between organisms that are related to each other?.4 How can comparing DNA sequences between different species provide information about evolution?.3 What is the reason for DNA similarity?.2 What does it indicate if two organisms have similar DNA sequences?.In practice, sequence evolution is mostly due to nucleotide mutations, deletions, and insertions (Figure 2.2). We must make some assumptions when performing sequence alignment, if only because we must transform a biological problem into a computationally feasible one and we require a model with relative simplicity and tractability. The goal of sequence alignment is to infer the ‘edit operations’ that change a genome by looking only at these endpoints. Thus, we are limited to comparing just the genomes of living descendants. Genomes change over time, and the scarcity of ancient genomes makes it virtually impossible to compare the genomes of living species with those of their ancestors. See Chapter ? for an in-depth discussion of evolutionary modeling and functional conservation in the context of genome annotation. By using known codon substitution frequencies and RNA secondary structure constraints, for example, we can calculate the probability that evolution acted to preserve a biological function. The most common approach to this problem involves modeling the evolutionary process. In order to extract accurate biological information from sequence alignments we have to separate true signatures from noise. We have to be cautious with our interpretations, however, because conservation does sometimes occur by random chance. 2 This example highlights how evolutionary data can help locate functional areas of the genome: per-nucleotide levels of conservation denote the importance of each nucleotide, and exons are among the most conserved elements in the genome. In particular, we note some small conserved motifs such as CGG and CGC, which in fact are functional elements in the binding of Gal4. As we look at this alignment, we note that some areas are more similar than others, suggesting that these areas have been conserved through evolution. As an example, we considered the alignment of the Gal10-Gal1 intergenic region for four different yeast species, the first cross-species whole genome alignment (Figure 2.1). These conserved regions typically imply functional elements and vice versa. Within orthologous gene sequences, there are islands of conservation, or relatively large stretches of nucleotides that are preserved between generations.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
